Comparing the end reason of reads in a nanopore experiment
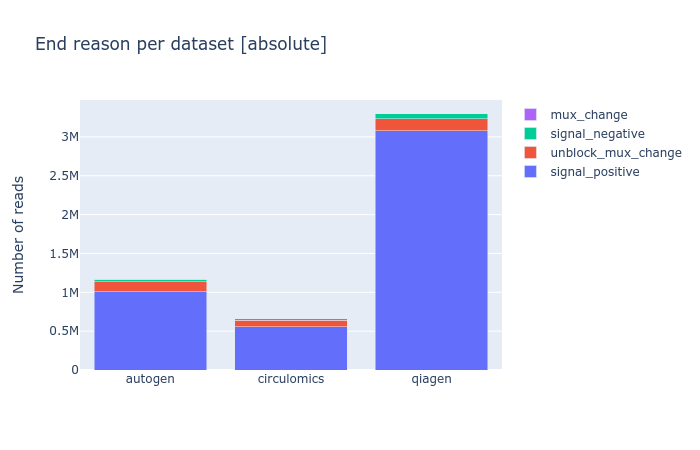
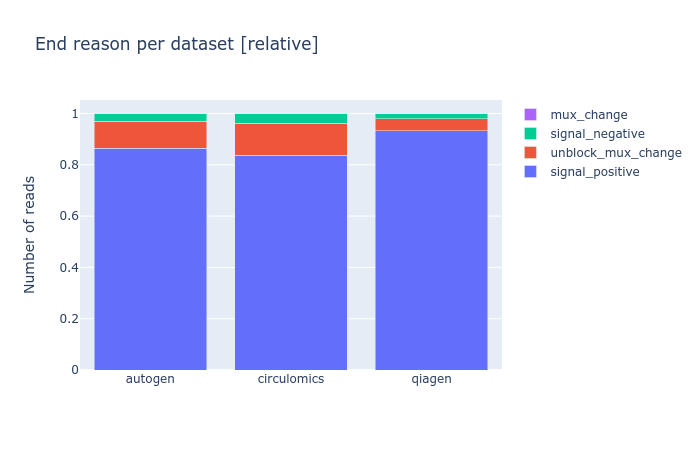
Since a recent version of MinKNOW, the software controlling a nanopore sequencer, a sequencing_summary file is created before basecalling, in which one column is of particular interest: the end_reason. Although I’m not yet sure what each value means, I believe it gives per read the reason why the software decided to stop sequencing here, mostly because the read was finished, but optionally because it was blocking the pore. Blocked pores is mainly an issue with very long reads, and it can drastically reduce the yield of your flow cell. So it’s something to an eye on.
I wrote a quick script to compare multiple of these pre-basecalling sequencing_summary files, available as extra script of my nanopore experiment comparison tool NanoComp at https://github.com/wdecoster/nanocomp/blob/master/scripts/end_reason_comparison.py. It will count the occurrences of all end_reason values, and show a stacked bar chart, both in absolute numbers and in relative fractions. More functionality here can be added, and contributions are definitely welcome.