Update on Oxford Nanopore basecall quality scores
Since an older post on basecall quality scores still attracts quite a lot of visits to my blog I think an update is timely.
In this post I look at a subset of PromethION data from NA19240, basecalled using Guppy 1.5.1 and aligned to the recommended set of GRCh38 using minimap2 v2.10. Using NanoPlot I generate the following plot, in which average basecall quality score is plotted against the percent reference identity or the edit distance scaled by read length, as a proxy for the accuracy.
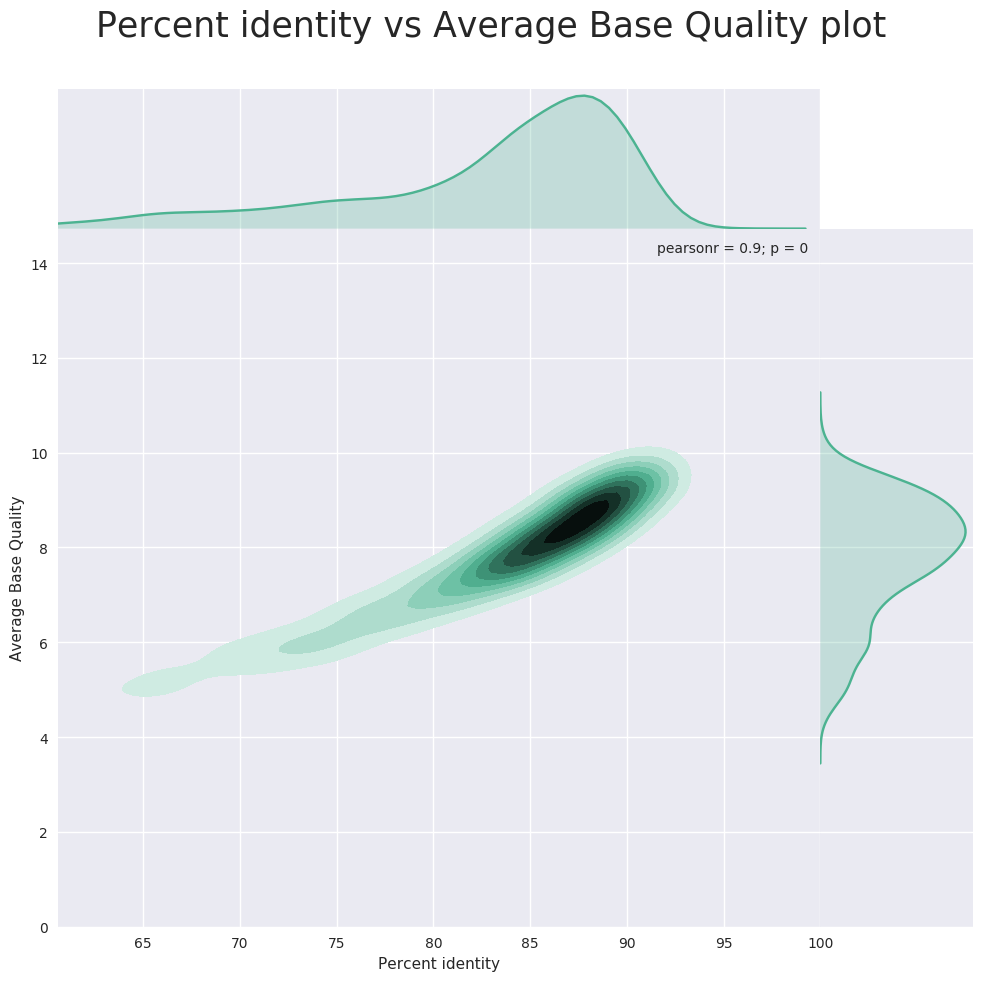 My conclusion from June 2017 was that the quality score was overconfident. The majority of reads can now be found at a quality score of ~8.5, which theoretically would correspond to an error rate of 14.1%. And that approximately corresponds to the percent identity, so it looks like the quality score is now well-calibrated!
My conclusion from June 2017 was that the quality score was overconfident. The majority of reads can now be found at a quality score of ~8.5, which theoretically would correspond to an error rate of 14.1%. And that approximately corresponds to the percent identity, so it looks like the quality score is now well-calibrated!