Inferring the sex based on transcriptome sequencing
Analogous to my previous post on inferring the sex of individuals based on exome sequencing I’ll now show you how to do the same for transcriptome sequencing. In the example, I use data from Lexogen QuantSeq but this is most likely equally applicable to other RNA-seq approaches. This is a useful QC step and can detect roughly 50% of sample swaps in your experiment.
This code is a function in my DEA.R R script for reproducible and convenient differential expression analysis from the command line. I use XIST as a female-specific gene and 4 chrY genes for male-specific expression (based on Staedtler et al 2013 ). It takes a vector of expected genders, a count matrix (e.g. from featureCounts) and a vector of sample names. The plot is made using ggplot2.
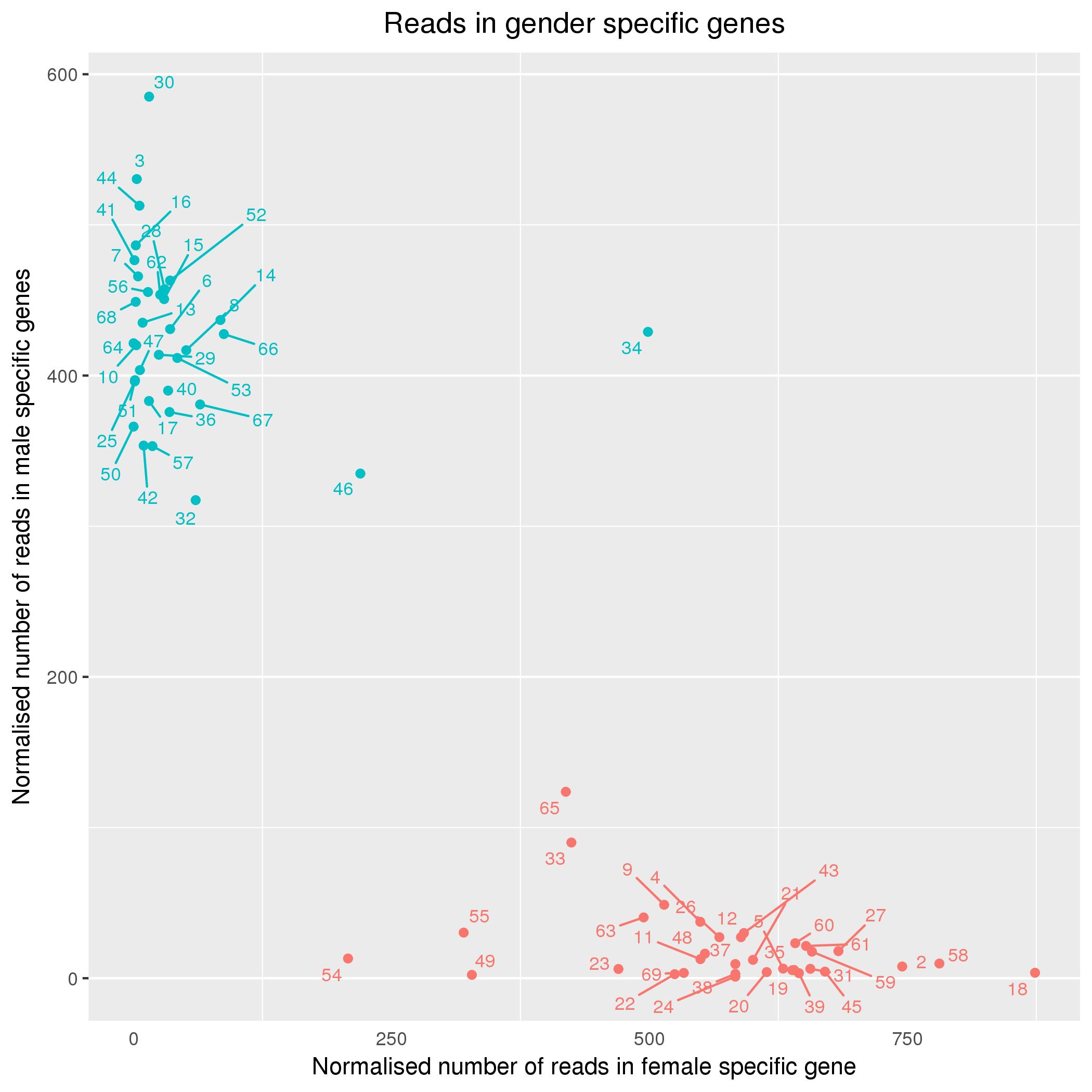
https://gist.github.com/wdecoster/561e4808a00011f141ce9ab32da3cb47
UPDATE: Devon Ryan suggested normalizing the read counts, of which I added the result (and the code) below. There is indeed some improvement, but not by a lot.
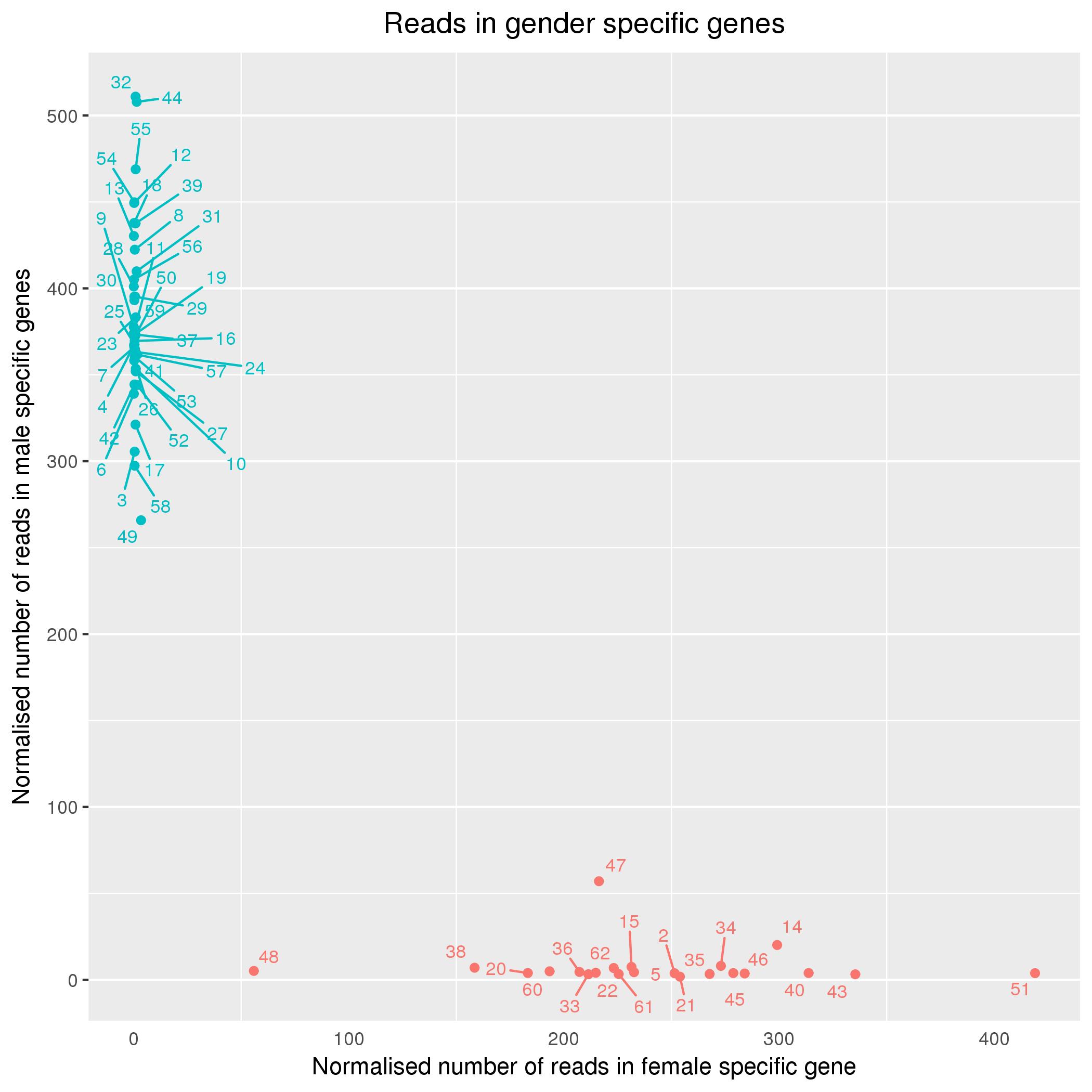
https://gist.github.com/wdecoster/eaa7fadbc23ce507a1ebe3d902c24eae